Documentation
Content:
- Summary of VIOLIN Technologies
- Data Curation Work Flow (also check: Data Curation Tutorial)
- Screenshots of VIOLIN components
- Database Schema
- Glossary (Click Here for direct access)
Summary of VIOLIN Technologies:
- Vaccine Ontology (VO): Dr. Yongqun "Oliver" He (VIOLIN) has initiated and led a community effort to develop the Vaccine Ontology (VO) for vaccine data standardization and analysis, literature mining, and automated reasoning. VO is being developed through collaborations with Dr. Barry Smith (University at Buffalo and the National Center for Biomedical Ontology), Dr. Lindsay Cowell (Duke University and the Infectious Disease Ontology), Dr. Harry Mobley (University of Michigan), and many other vaccine and ontology experts.
- Vaccine literature mining: VIOLIN provide several literature mining tools, including natural language processing (NLP)-based Vaxpresso, MeSH-based Vaxmesh, Vaccine alert system Vaxlert, and general vaccine literature search Litesearch. Vomine will become a vaccine-specific natural language processing (NLP)-based literature mining program using the Vaccine Ontology (VO), Gene Ontology (GO), Medical Subject Headings (MeSH), and other ontology or controlled vocabulary systems. Vomine will be developed by VIOLIN through collaboration with the National Center for Integrative Biomedical Informatics (NCIBI).
- Vaccine & vaccine components: Manually curated vaccine data can be queried and analyzed using our Vaxquery search engine. Vaccine-related genes can be queried with Vaxgen and analyzed using a customized BLAST search program. We are also including information on vaccine adjuvants (Vaxjo) and vaccine vectors (Vaxvec).
- Vaccine Mechanisms: The mechanisms of how vaccines against various diseases are the topics of Vaxism. Vaxmod specfically targets to study different animal models used in vaccine R & D. We also plan to build a Vaxnet program which will analyze vaccine-induced response networks using ontology-based information integration, NLP-based literature mining, and other statistical techniques.
 Vaxign (beta): A new vaccine target design program is available for public use. Vaxign predicts all possible vaccine tagets based on pathogen genomes or used provided sequences. It is an ideal tool for reverse vaccinology study.
Vaxign (beta): A new vaccine target design program is available for public use. Vaxign predicts all possible vaccine tagets based on pathogen genomes or used provided sequences. It is an ideal tool for reverse vaccinology study.- Vaccine Society: We have build several web programs to cover various aspects of vaccine society, including vaccine R & D experts (Vaxperts), vaccine information targeting the public (VaxPub), vaccine companies (VaxCom), vaccine-related regulations ns (VaxLaw), vaccine media (VaxMedia), vaccine meetings (VaxMeet), vaccine funding and grants (VaxFund), vaccine R & D career opportunties (VaxCareer), and a wiki system for vaccine R & D (Vaccine Wiki).
- Data Download and Exchange: The curated vaccine data in VIOLIN can be downloaded through an XML-based VIOLIN data format VIOLINML or via a web service called V-Utilities.
Data Curation Work Flow:
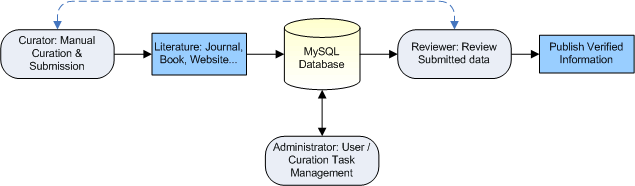
NOTE: The VIOLIN data submission model is similar to the GeneRIF model that is used by NCBI Entrez Gene to submit the functional annotation of genes.
Link To: Data Curation Tutorial
Screenshots of VIOLIN components
We have prepared a few screenshots for explanation of some VIOLIN components.
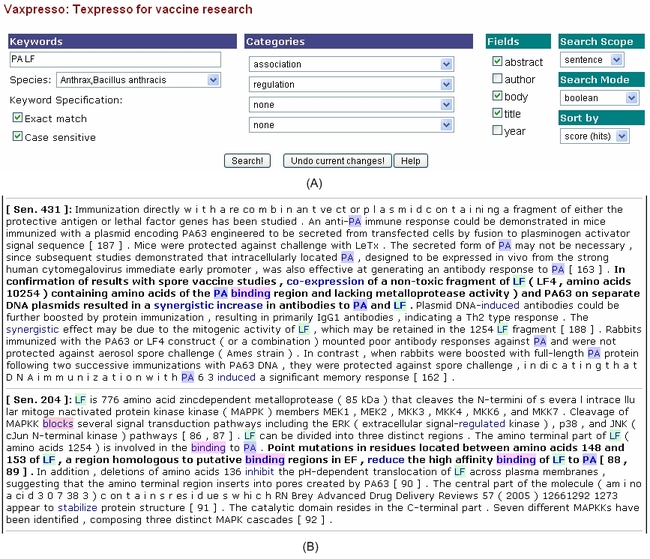
(1). Vaxpresso literature search. This example uses a natural language processing (NLP)-based information retrieval program to search for the interaction between PA (Protective Antigen) and LF (Lethal Factor) in Bacillus anthracis (anthrax) (A). This query resulted in 73 matches in 39 publications (data not shown). The keywords PA, LF and any word under the “association” or “regulation” category (e.g., co-expression, binding, induced, increase) are highlighted (B). Sentences containing both keywords and a categorized word are shown in bold typeface. It is highly suggested that PA and LF used together will aide in developing a better vaccine and that a given region of LF is critical for binding to EF (Edema Factor) and PA.

(2). A Vaxmesh search for keyword “interferon”. All vaccine related literature publications stored in VIOLIN can be visualized by the interactive MeSH-based tree browser. The two hotlinked numbers in each line link to all publications containing the term as a MeSH term or a major end-node MeSH term, respectively. Keywords can be further searched for more specific results. The figure shows the hierarchical MeSH tree structure leading to different topics under Gene Expression Regulation related to the keyword “interferon”.
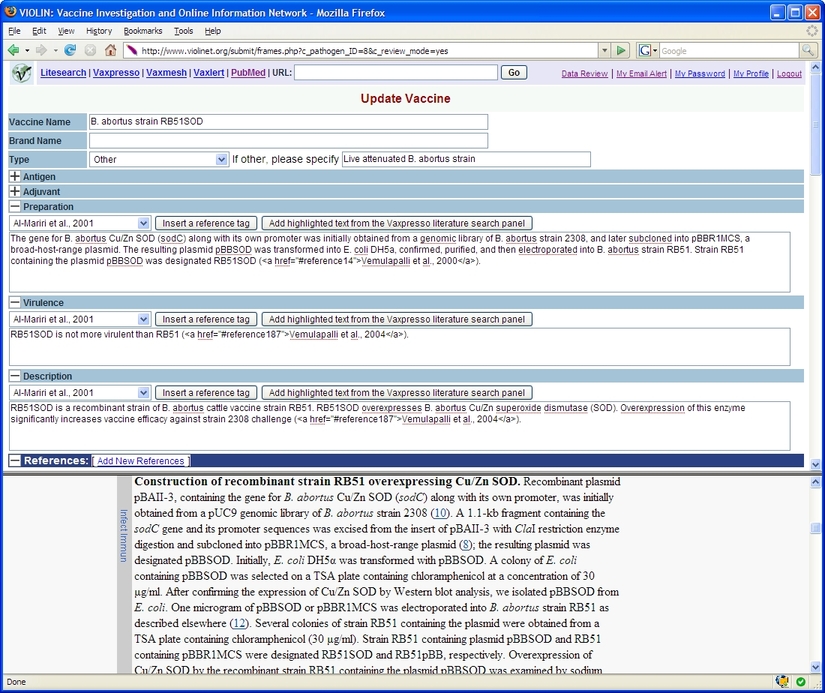
(3). VIOLIN data submission system. The system is based on Limix, an interactive system that integrates computational literature mining and manual curation. The top manual curation frame contains structured editable text fields for a curator to copy, edit, and submit updated vaccine information to a backend database. Different sections of information can be folded for convenient visualization and space saving. The top frame also allows a curator to choose an internal literature search program, search PubMed, or type a URL to access a web site. The results are shown in the bottom text mining frame. A curator can easily copy text from the bottom frame to the editable text fields in the top frame. The horizontal line between two frames can be adjusted to efficiently distribute the space. The bottom frame in this figure shows part of the full text of a peer-reviewed manuscript available in PubMed Central.

(4). Curated vaccine comparison. This figure shows comparison of two vaccines involved in “superoxide dismutase”. These two vaccines can be shown up during a keyword search of “superoxide dismutase” in Vaxquery.
Database schema:
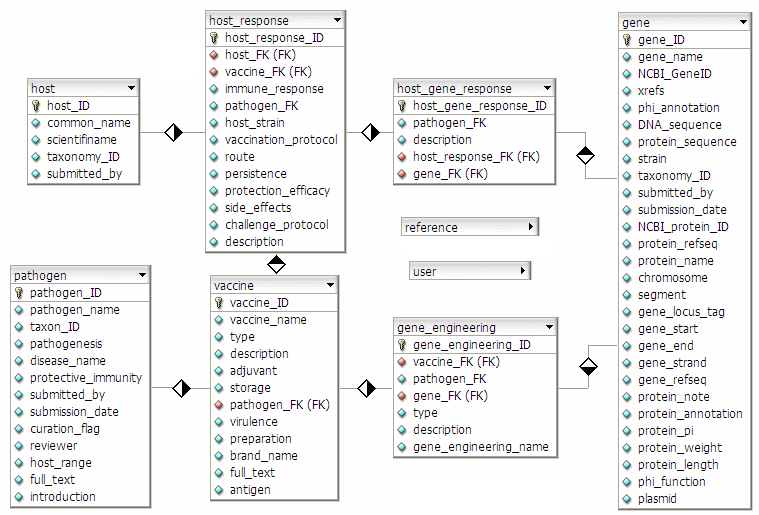
Glossary
We have collected vaccine-related definitions for users' reference. Please click Here.
